2026-04-16
•Tobia Ochsner
•3 min read
A First Look at the BEAST 2.8 Parser
The first version of a proper PhyloSpec parser for BEAST 2.8 is done 🎉 (PR). This post walks through its main features and what is still to come.
The parser takes a PhyloSpec model, runs the static type checker and linter to catch common errors, and — if the model is valid — constructs a full BEAST 2 analysis ready to run. BEAST then takes over and performs the MCMC.
An example of a model which can be run is the following:
Alignment data = fromNexus("primate-mtDNA.nex")
Alignment filtered = subset(
alignment=data,
start=10,
end=898
)
Tree tree ~ Yule(
birthRate~LogNormal(mean=1.0, logSd=2.0),
taxa=taxa(filtered)
)
QMatrix qMatrix = hky(
kappa~LogNormal(mean=1, logSd=1.0),
baseFrequencies=[0.25, 0.25, 0.25, 0.25]
)
Vector<Rate> branchRates ~ StrictClock(
clockRate~LogNormal(mean=0.1, logSd=2.0),
tree,
)
Vector<Rate> siteRates ~ DiscreteGammaInv(
shape~LogNormal(mean=0.1, logSd=2.0),
numCategories=4,
invariantProportion=0.1,
numSites=numSites(filtered)
)
Alignment alignment ~ PhyloCTMC(
tree, qMatrix, branchRates, siteRates
) observed as filtered
Age root = rootAge(tree) observed between [0.01, 3.0]
mcmc {
Integer chainLength = 100000
Logger treeLogger = treeLogger(
file="trees.trees", logEvery=1000
)
}Supported PhyloSpec Features
48 out of 76 core components are at least partially supported (see the current status for the full picture). Most of the gaps — mk, PhyloOU, and similar — have no equivalent in BEAST core and will only be unlocked once external package support is added.
A handful of components are only partially supported. exp(x), for instance, works only when x is a constant or deterministic expression, or when used inside a logger. There is simply no BEAST StateNode that implements the exponential function.
On the language side, indexed statements (like Real x[i] = f(i) for i in 1:10), matrix literals, and index accessors are not yet handled.
Flexibility
Internally, the parser uses a flexible pattern-matching framework, so the same model can be written in many equivalent ways. The following three snippets all produce identical BEAST objects:
Alignment alignment ~ PhyloCTMC(
tree,
qMatrix,
branchRates~StrictClock(
clockRate~LogNormal(mean=1.0, logSd=0.5), tree
)
) observed as dataRate clockRate ~ LogNormal(mean=1.0, logSd=0.5)
Vector<Rate> branchRates ~ StrictClock(clockRate, tree)
Alignment alignment ~ PhyloCTMC(
tree, qMatrix, branchRates
) observed as dataRate clockRate ~ LogNormal(logMean=log(1.0), logSd=0.5)
Rate branchRates[i] = clockRate for i in 1:numBranches(tree)
Alignment alignment ~ PhyloCTMC(
tree, qMatrix, branchRates
) observed as dataLoggers
By default, the parser creates a screen logger, a file logger, and a tree logger. Every named variable and every inline random variable is included automatically:
Rate birthRate ~ LogNormal(logMean=1.0, logSd=0.5)
Real logBirthRate = log(birthRate)
Tree tree ~ Yule(
birthRate, tree
)
Real halfRootAge = rootAge(tree) / 2
// birthRate, logBirthRate, tree, and halfRootAge are loggedTo override the defaults, use the mcmc block:
mcmc {
Logger screenLogger = screenLogger(
logEvery=100
)
Logger fileLogger = fileLogger(
file="log.log", logEvery=1000, parameters=[birthRate]
)
Logger treeLogger = treeLogger(
file="trees.trees", logEvery=10000
)
}
Operator Selection
Operators are selected automatically based on the state node types. A few multi-variable rules are also applied — for example, an UpDownOperator is added whenever there is both a tree and a clock rate coming together in a PhyloCMTC distribution.
A future version will allow operators to be configured explicitly via a beast28 block.
Errors
A lot of effort went into making error messages actionable. Here are a few examples:
Wrong argument names:

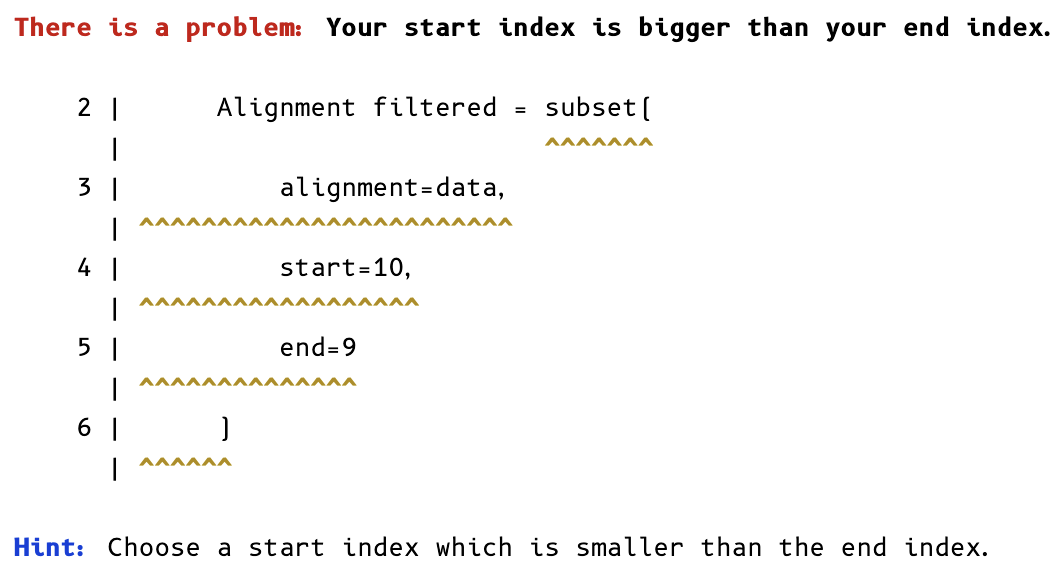
Invalid values:
Unsupported statements:
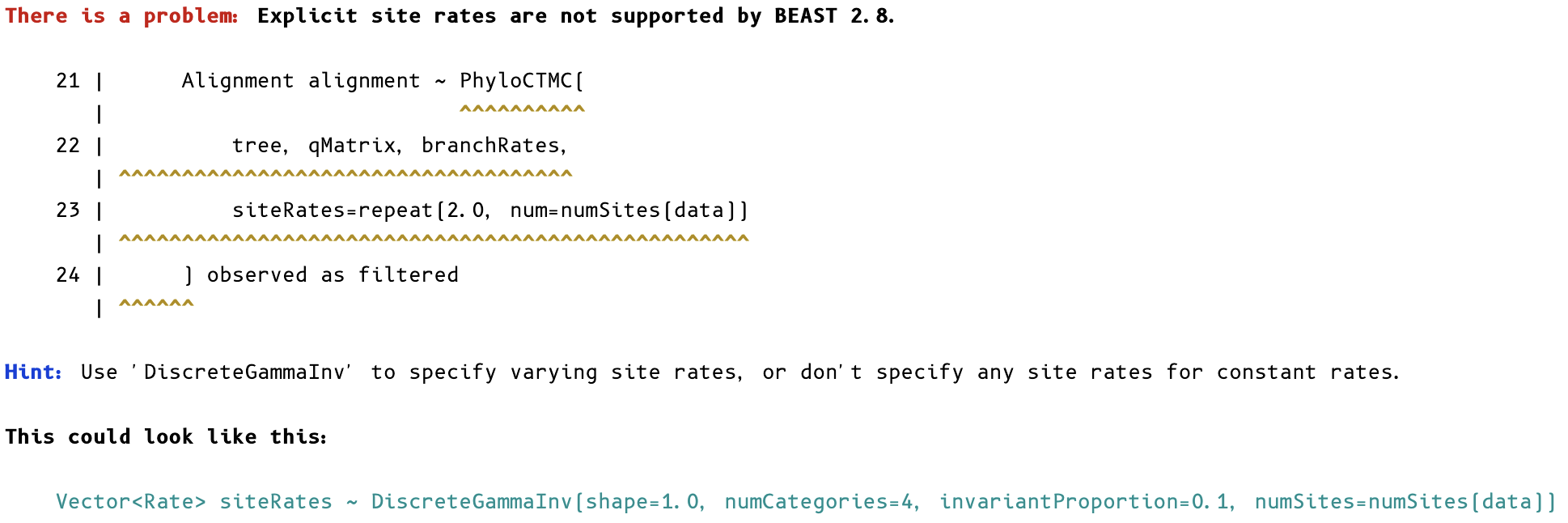
Instantiation errors:
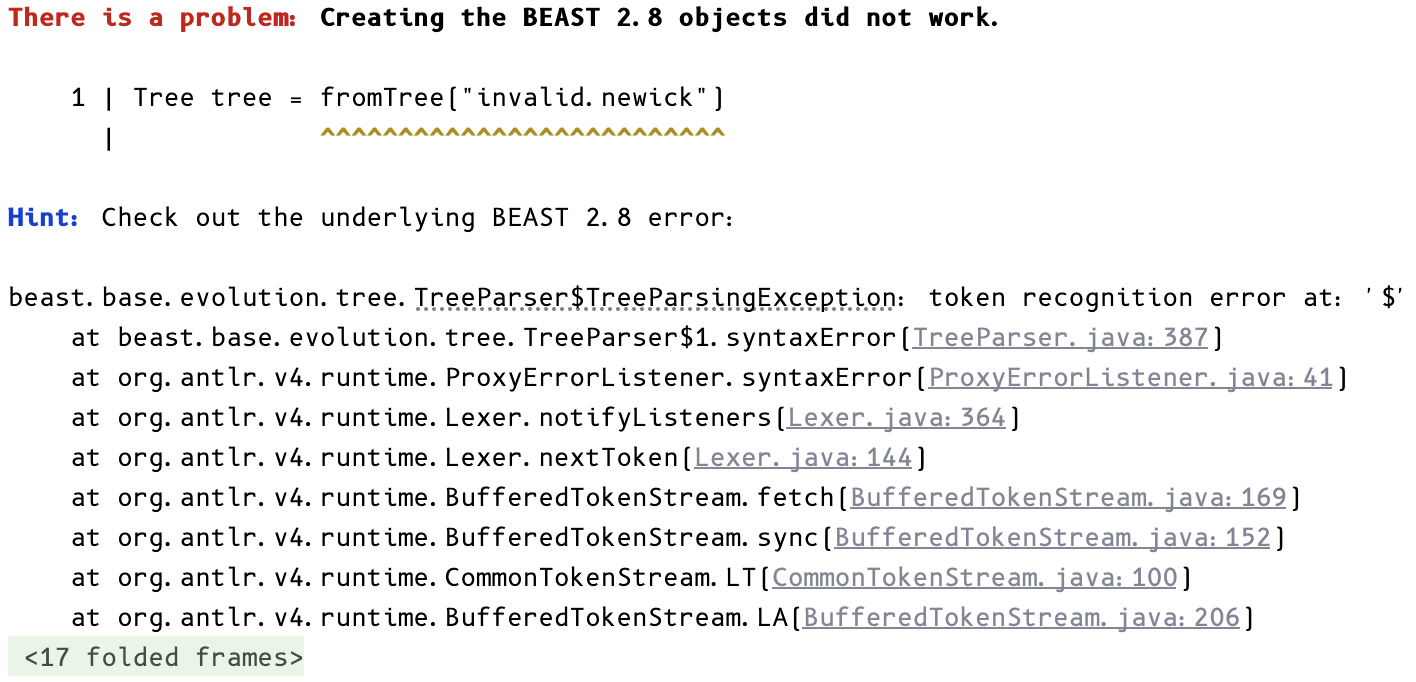
In the last example, invalid.newick does not contain a valid tree. When BEAST itself raises an error, the parser traces it back to the relevant PhyloSpec lines while still surfacing the underlying message.
As a side effect, precise errors also make the parser much more useful when building models with an AI agent.
Implementation
A follow-up post will go into the algorithm in detail. In short, the parser is built from over 70 small Java classes called tiles, each responsible for mapping one fragment of a PhyloSpec model onto the corresponding BEAST objects.
As a concrete example, there are two tiles for the exp function. One wraps BEAST’s RPNCalculator and is used when the result needs to be logged. The other handles any non-stochastic input more directly:
public class ExpTile extends GeneratorTile<RealScalarParam<PositiveReal>> {
@Override
public String getPhyloSpecGeneratorName() {
return "exp";
}
GeneratorTileInput<RealScalarParam<? extends Real>> xInput = new GeneratorTileInput<>(
"x", Set.of(Stochasticity.CONSTANT, Stochasticity.DETERMINISTIC)
);
@Override
public RealScalarParam<PositiveReal> applyTile(BEASTState beastState) {
RealScalarParam<? extends Real> variable = this.xInput.apply(beastState);
return new RealScalarParam<>(Math.exp(variable.get()), PositiveReal.INSTANCE);
}
}The algorithm automatically selects and assembles the right tiles for any given model.
Next Steps
One important addition is external package support, which would immediately unlock components like FossilizedBirthDeath and mk. Other planned features include a beast28 block for configuring operators directly, filling in the remaining language gaps (indexed statements, matrix literals), and building a validation suite that compares the constructed BEAST DAG against hand-crafted XMLs. The parser will also be integrated into phylorun, making it possible to run PhyloSpec models from the command line without any manual setup.